| PRKAA1 |
|---|
 |
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
4RED, 4RER, 4REW, 5EZV |
|
|
| Identifiers |
|---|
| Aliases | PRKAA1, AMPK, AMPKa1, Protein kinase, AMP-activated, alpha 1, protein kinase AMP-activated catalytic subunit alpha 1 |
|---|
| External IDs | OMIM: 602739; MGI: 2145955; HomoloGene: 49590; GeneCards: PRKAA1; OMA:PRKAA1 - orthologs |
|---|
| EC number | 2.7.11.26 |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 5 (human)[1] |
|---|
| | Band | n/a | Start | 40,759,379 bp[1] |
|---|
| End | 40,798,374 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 15 (mouse)[2] |
|---|
| | Band | 15|15 A1 | Start | 5,173,343 bp[2] |
|---|
| End | 5,211,380 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - Achilles tendon
- sperm
- rectum
- gallbladder
- gastric mucosa
- stromal cell of endometrium
- epithelium of colon
- tibial arteries
- body of pancreas
- Descending thoracic aorta
|
| | Top expressed in | - granulocyte
- islet of Langerhans
- epithelium of small intestine
- mammillary body
- white adipose tissue
- hand
- ventromedial nucleus
- medial vestibular nucleus
- lateral hypothalamus
- ventral tegmental area
|
| | More reference expression data |
|
|---|
| BioGPS | 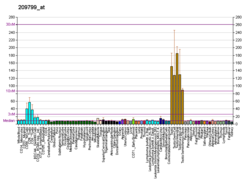 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - transferase activity
- nucleotide binding
- protein kinase activity
- cAMP-dependent protein kinase activity
- [acetyl-CoA carboxylase kinase activity]
- AMP-activated protein kinase activity
- histone serine kinase activity
- chromatin binding
- metal ion binding
- kinase activity
- protein serine/threonine kinase activity
- tau-protein kinase activity
- protein C-terminus binding
- protein binding
- [hydroxymethylglutaryl-CoA reductase (NADPH) kinase activity]
- ATP binding
- tau protein binding
| | Cellular component | - nucleotide-activated protein kinase complex
- apical plasma membrane
- nucleoplasm
- nucleus
- cytoplasm
- cytosol
- nuclear speck
- intracellular anatomical structure
- axon
- dendrite
- protein-containing complex
- neuronal cell body
| | Biological process | - cellular response to organonitrogen compound
- steroid metabolic process
- cellular response to glucose starvation
- sterol biosynthetic process
- lipid biosynthetic process
- regulation of transcription, DNA-templated
- response to hypoxia
- cellular response to ethanol
- glucose homeostasis
- positive regulation of skeletal muscle tissue development
- protein heterooligomerization
- rhythmic process
- phosphorylation
- lipid metabolism
- regulation of vesicle-mediated transport
- negative regulation of apoptotic process
- cholesterol metabolic process
- Wnt signaling pathway
- response to activity
- fatty acid metabolic process
- transcription, DNA-templated
- cellular response to prostaglandin E stimulus
- autophagy
- protein phosphorylation
- negative regulation of TOR signaling
- cold acclimation
- positive regulation of gene expression
- response to camptothecin
- fatty acid biosynthetic process
- fatty acid homeostasis
- regulation of circadian rhythm
- fatty acid oxidation
- regulation of peptidyl-serine phosphorylation
- cellular response to nutrient levels
- positive regulation of cell population proliferation
- negative regulation of glucosylceramide biosynthetic process
- glucose metabolic process
- positive regulation of cholesterol biosynthetic process
- response to caffeine
- cholesterol biosynthetic process
- negative regulation of lipid catabolic process
- response to gamma radiation
- macroautophagy
- response to UV
- positive regulation of glycolytic process
- response to hydrogen peroxide
- cellular response to hypoxia
- signal transduction
- cellular response to hydrogen peroxide
- steroid biosynthetic process
- regulation of signal transduction by p53 class mediator
- positive regulation of autophagy
- CAMKK-AMPK signaling cascade
- regulation of macroautophagy
- chromatin organization
- positive regulation of mitochondrial transcription
- positive regulation of protein targeting to mitochondrion
- negative regulation of gene expression
- cellular response to oxidative stress
- intracellular signal transduction
- negative regulation of insulin receptor signaling pathway
- motor behavior
- regulation of stress granule assembly
- neuron cellular homeostasis
- regulation of microtubule cytoskeleton organization
- cellular response to calcium ion
- cellular response to glucose stimulus
- energy homeostasis
- positive regulation of protein localization
- negative regulation of tubulin deacetylation
- positive regulation of peptidyl-lysine acetylation
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | | |
|---|
| RefSeq (protein) | NP_006242
NP_996790
NP_001341957
NP_001341958
NP_001341963
|
|---|
NP_001341964
NP_001341965
NP_001341966 |
| |
|---|
| Location (UCSC) | Chr 5: 40.76 – 40.8 Mb | Chr 15: 5.17 – 5.21 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|

 2h6d: Protein Kinase Domain of the Human 5'-AMP-activated protein kinase catalytic subunit alpha-2 (AMPK alpha-2 chain)
2h6d: Protein Kinase Domain of the Human 5'-AMP-activated protein kinase catalytic subunit alpha-2 (AMPK alpha-2 chain)

















